-MEDIATED_SIGNALING_105.png)
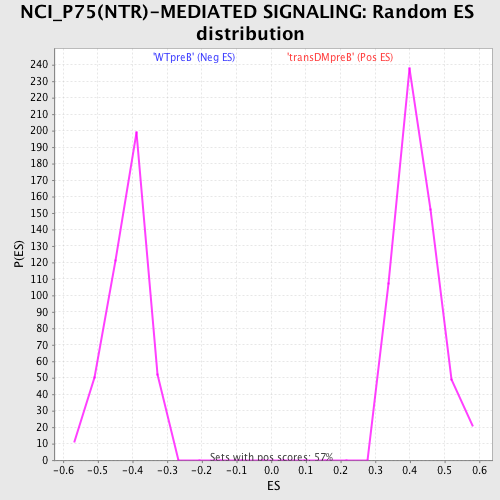
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | NCI_P75(NTR)-MEDIATED SIGNALING |
| Enrichment Score (ES) | -0.64293003 |
| Normalized Enrichment Score (NES) | -1.5495468 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1223675 |
| FWER p-Value | 0.809 |
-MEDIATED_SIGNALING_105.png)
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CCND1 | 4487 4488 8707 17535 | 85 | 2.719 | 0.0358 | No | ||
| 2 | TIAM1 | 22547 | 687 | 1.091 | 0.0195 | No | ||
| 3 | YWHAQ | 10369 | 1061 | 0.761 | 0.0106 | No | ||
| 4 | CYCS | 8821 | 1112 | 0.728 | 0.0187 | No | ||
| 5 | MAGEH1 | 24029 | 1207 | 0.670 | 0.0236 | No | ||
| 6 | ARRB2 | 20806 | 1303 | 0.619 | 0.0277 | No | ||
| 7 | IRAK1 | 4916 | 1367 | 0.587 | 0.0330 | No | ||
| 8 | SRF | 22961 1597 | 1447 | 0.548 | 0.0368 | No | ||
| 9 | MALT1 | 6274 | 1542 | 0.516 | 0.0394 | No | ||
| 10 | YWHAZ | 10370 | 1567 | 0.504 | 0.0456 | No | ||
| 11 | NFKB2 | 23810 | 1631 | 0.479 | 0.0493 | No | ||
| 12 | FAIM | 19344 | 1717 | 0.449 | 0.0514 | No | ||
| 13 | IKBKB | 4907 | 1740 | 0.441 | 0.0568 | No | ||
| 14 | CHUK | 23665 | 1827 | 0.403 | 0.0581 | No | ||
| 15 | PRKCZ | 5260 | 1997 | 0.346 | 0.0541 | No | ||
| 16 | YWHAE | 20776 | 2133 | 0.303 | 0.0513 | No | ||
| 17 | E2F1 | 14384 | 2208 | 0.278 | 0.0514 | No | ||
| 18 | MAPK8 | 6459 | 2251 | 0.267 | 0.0531 | No | ||
| 19 | RPS6KA1 | 15725 | 2384 | 0.231 | 0.0494 | No | ||
| 20 | MYD88 | 18970 | 2481 | 0.202 | 0.0472 | No | ||
| 21 | MAP2K1 | 19082 | 2640 | 0.166 | 0.0411 | No | ||
| 22 | MEF2C | 3204 9378 | 2762 | 0.145 | 0.0367 | No | ||
| 23 | DIABLO | 16375 | 2875 | 0.125 | 0.0325 | No | ||
| 24 | APP | 4402 | 2989 | 0.108 | 0.0280 | No | ||
| 25 | CDK5R1 | 20745 | 2995 | 0.108 | 0.0293 | No | ||
| 26 | CASP3 | 8693 | 3119 | 0.092 | 0.0240 | No | ||
| 27 | PLG | 1593 23383 | 3208 | 0.082 | 0.0205 | No | ||
| 28 | RTN4 | 7569 1328 1343 1442 | 3265 | 0.076 | 0.0186 | No | ||
| 29 | DNAJA3 | 1732 13518 | 3357 | 0.070 | 0.0147 | No | ||
| 30 | BCL2L11 | 2790 14861 | 3579 | 0.055 | 0.0035 | No | ||
| 31 | ARHGDIA | 20116 | 3705 | 0.049 | -0.0025 | No | ||
| 32 | GIPC1 | 3841 3882 18552 | 3828 | 0.044 | -0.0085 | No | ||
| 33 | RAPGEF1 | 4218 2860 | 3945 | 0.040 | -0.0142 | No | ||
| 34 | AKT1 | 8568 | 4202 | 0.032 | -0.0276 | No | ||
| 35 | FOS | 21202 | 4235 | 0.031 | -0.0288 | No | ||
| 36 | ABL1 | 2693 4301 2794 | 4331 | 0.029 | -0.0335 | No | ||
| 37 | SHC1 | 9813 9812 5430 | 4437 | 0.027 | -0.0388 | No | ||
| 38 | STAT5A | 20664 | 4526 | 0.025 | -0.0432 | No | ||
| 39 | NTRK2 | 9491 | 4545 | 0.025 | -0.0438 | No | ||
| 40 | NTRK3 | 2069 3652 3705 9492 | 4552 | 0.025 | -0.0438 | No | ||
| 41 | CAMK2A | 2024 23541 1980 | 4715 | 0.022 | -0.0522 | No | ||
| 42 | HRAS | 4868 | 4722 | 0.022 | -0.0522 | No | ||
| 43 | FRS3 | 23209 | 4728 | 0.022 | -0.0522 | No | ||
| 44 | NTF3 | 16991 | 4821 | 0.021 | -0.0568 | No | ||
| 45 | NDN | 18222 | 5204 | 0.017 | -0.0773 | No | ||
| 46 | IKBKG | 2570 2562 4908 | 5311 | 0.016 | -0.0828 | No | ||
| 47 | RASGRF1 | 3050 19357 | 5369 | 0.015 | -0.0857 | No | ||
| 48 | MATK | 3425 19930 | 5426 | 0.015 | -0.0885 | No | ||
| 49 | SHC3 | 21465 | 5554 | 0.014 | -0.0952 | No | ||
| 50 | RIPK2 | 2528 15935 | 5631 | 0.013 | -0.0991 | No | ||
| 51 | APH1A | 15501 5947 10378 | 5915 | 0.011 | -0.1142 | No | ||
| 52 | MAG | 17880 | 5926 | 0.011 | -0.1146 | No | ||
| 53 | FBXW11 | 20926 | 5928 | 0.011 | -0.1145 | No | ||
| 54 | BDNF | 14926 2797 | 6229 | 0.010 | -0.1306 | No | ||
| 55 | MMP7 | 19568 | 6250 | 0.010 | -0.1315 | No | ||
| 56 | SRC | 5507 | 6276 | 0.009 | -0.1328 | No | ||
| 57 | RHOA | 8624 4409 4410 | 6499 | 0.009 | -0.1447 | No | ||
| 58 | RAP1A | 8467 | 7144 | 0.006 | -0.1795 | No | ||
| 59 | CDC42 | 4503 8722 4504 2465 | 7364 | 0.005 | -0.1912 | No | ||
| 60 | RHOC | 15467 | 7450 | 0.005 | -0.1958 | No | ||
| 61 | SFN | 7060 12062 | 7743 | 0.004 | -0.2115 | No | ||
| 62 | RIT2 | 5380 | 8134 | 0.003 | -0.2326 | No | ||
| 63 | XPO1 | 4172 | 8147 | 0.003 | -0.2332 | No | ||
| 64 | MMP3 | 19572 3148 | 8689 | 0.002 | -0.2625 | No | ||
| 65 | ATM | 2976 19115 | 9162 | 0.001 | -0.2880 | No | ||
| 66 | DNM1 | 2777 8857 | 9407 | 0.000 | -0.3012 | No | ||
| 67 | PRKCI | 9576 | 9472 | 0.000 | -0.3047 | No | ||
| 68 | OMG | 5209 | 9691 | -0.001 | -0.3165 | No | ||
| 69 | PLCG1 | 14753 | 9774 | -0.001 | -0.3209 | No | ||
| 70 | PSEN1 | 5297 2125 9630 | 10258 | -0.002 | -0.3471 | No | ||
| 71 | NRAS | 5191 | 10314 | -0.002 | -0.3500 | No | ||
| 72 | RPS6KA5 | 2076 21005 | 10489 | -0.002 | -0.3594 | No | ||
| 73 | RTN4R | 22830 | 10611 | -0.003 | -0.3659 | No | ||
| 74 | YWHAH | 5937 10368 | 10669 | -0.003 | -0.3690 | No | ||
| 75 | SOS1 | 5476 | 10751 | -0.003 | -0.3733 | No | ||
| 76 | CAMK4 | 4473 | 10887 | -0.004 | -0.3806 | No | ||
| 77 | PIK3R1 | 3170 | 11690 | -0.006 | -0.4239 | No | ||
| 78 | NTRK1 | 15299 | 12042 | -0.007 | -0.4428 | No | ||
| 79 | PTPN11 | 5326 16391 9660 | 12084 | -0.007 | -0.4449 | No | ||
| 80 | CREB1 | 3990 8782 4558 4093 | 12093 | -0.007 | -0.4453 | No | ||
| 81 | TRPC3 | 15355 | 12642 | -0.010 | -0.4748 | No | ||
| 82 | TNF | 23004 | 12661 | -0.010 | -0.4756 | No | ||
| 83 | NGFR | 5174 | 13147 | -0.013 | -0.5017 | No | ||
| 84 | CASP6 | 15422 1885 | 13205 | -0.013 | -0.5046 | No | ||
| 85 | FOXO3 | 19782 3402 | 13423 | -0.015 | -0.5161 | No | ||
| 86 | SORT1 | 15452 5475 9847 | 13489 | -0.016 | -0.5194 | No | ||
| 87 | MAP3K2 | 11165 | 13691 | -0.018 | -0.5301 | No | ||
| 88 | CRK | 4559 1249 | 13772 | -0.019 | -0.5341 | No | ||
| 89 | PIK3CA | 9562 | 13925 | -0.021 | -0.5420 | No | ||
| 90 | GAB2 | 1821 18184 2025 | 14144 | -0.024 | -0.5535 | No | ||
| 91 | ERC1 | 1013 17021 995 1136 | 14333 | -0.027 | -0.5633 | No | ||
| 92 | MAGED1 | 24108 | 14402 | -0.029 | -0.5665 | No | ||
| 93 | MCF2L | 18680 5082 | 14495 | -0.030 | -0.5711 | No | ||
| 94 | RUSC1 | 1853 15283 | 14532 | -0.031 | -0.5725 | No | ||
| 95 | MAPK10 | 11169 | 14546 | -0.032 | -0.5728 | No | ||
| 96 | MAPK7 | 1381 20414 | 15001 | -0.047 | -0.5967 | No | ||
| 97 | TNFRSF1A | 1181 10206 | 15227 | -0.057 | -0.6080 | No | ||
| 98 | NOD2 | 6384 | 15341 | -0.064 | -0.6132 | No | ||
| 99 | BCL3 | 8654 | 15460 | -0.074 | -0.6185 | No | ||
| 100 | MAP2K5 | 19088 | 15507 | -0.079 | -0.6198 | No | ||
| 101 | CASP9 | 16001 2410 2458 | 15518 | -0.080 | -0.6191 | No | ||
| 102 | FURIN | 9538 | 15674 | -0.093 | -0.6261 | No | ||
| 103 | YWHAB | 14744 | 15786 | -0.105 | -0.6306 | No | ||
| 104 | NGFRAP1 | 24246 | 15886 | -0.118 | -0.6342 | No | ||
| 105 | RGS19 | 14303 | 15983 | -0.137 | -0.6374 | No | ||
| 106 | MAP3K14 | 11998 | 16026 | -0.144 | -0.6375 | No | ||
| 107 | CDK5 | 16591 | 16077 | -0.157 | -0.6379 | No | ||
| 108 | REL | 9716 | 16087 | -0.160 | -0.6360 | No | ||
| 109 | MAP2K3 | 20856 | 16216 | -0.188 | -0.6401 | Yes | ||
| 110 | YWHAG | 16339 | 16238 | -0.194 | -0.6384 | Yes | ||
| 111 | RASA1 | 10174 | 16249 | -0.195 | -0.6360 | Yes | ||
| 112 | MAPK14 | 23313 | 16271 | -0.200 | -0.6342 | Yes | ||
| 113 | NDNL2 | 17804 | 16273 | -0.200 | -0.6313 | Yes | ||
| 114 | CYLD | 18532 | 16280 | -0.201 | -0.6286 | Yes | ||
| 115 | BIRC3 | 19247 | 16305 | -0.207 | -0.6268 | Yes | ||
| 116 | RELB | 17942 | 16324 | -0.211 | -0.6247 | Yes | ||
| 117 | PRKCD | 21897 | 16416 | -0.233 | -0.6261 | Yes | ||
| 118 | RAF1 | 17035 | 16474 | -0.248 | -0.6255 | Yes | ||
| 119 | APAF1 | 8606 | 16701 | -0.317 | -0.6331 | Yes | ||
| 120 | UBE2D3 | 7253 | 16771 | -0.341 | -0.6317 | Yes | ||
| 121 | ADAM17 | 4343 | 16871 | -0.384 | -0.6314 | Yes | ||
| 122 | GRB2 | 20149 | 16888 | -0.389 | -0.6265 | Yes | ||
| 123 | SMPD2 | 5462 | 16959 | -0.418 | -0.6240 | Yes | ||
| 124 | MAP2K6 | 20614 1414 | 16980 | -0.426 | -0.6188 | Yes | ||
| 125 | MAPK3 | 6458 11170 | 17011 | -0.437 | -0.6139 | Yes | ||
| 126 | RAC1 | 16302 | 17052 | -0.451 | -0.6094 | Yes | ||
| 127 | BIRC2 | 4397 4398 | 17138 | -0.482 | -0.6068 | Yes | ||
| 128 | PRKCA | 20174 | 17237 | -0.520 | -0.6044 | Yes | ||
| 129 | GSK3B | 22761 | 17503 | -0.658 | -0.6089 | Yes | ||
| 130 | NCSTN | 930 12236 | 17510 | -0.662 | -0.5994 | Yes | ||
| 131 | SYK | 21636 | 17551 | -0.690 | -0.5913 | Yes | ||
| 132 | RELA | 23783 | 17583 | -0.705 | -0.5825 | Yes | ||
| 133 | BAD | 24000 | 17587 | -0.709 | -0.5721 | Yes | ||
| 134 | NFKB1 | 15160 | 17682 | -0.778 | -0.5657 | Yes | ||
| 135 | CRKL | 4560 | 17704 | -0.794 | -0.5550 | Yes | ||
| 136 | RAP1B | 10083 | 17748 | -0.823 | -0.5451 | Yes | ||
| 137 | NEDD4L | 8192 | 17892 | -0.942 | -0.5388 | Yes | ||
| 138 | NFKBIA | 21065 | 18048 | -1.142 | -0.5302 | Yes | ||
| 139 | BCL10 | 15397 | 18079 | -1.187 | -0.5142 | Yes | ||
| 140 | PDPK1 | 23097 | 18082 | -1.189 | -0.4966 | Yes | ||
| 141 | TRAF6 | 5797 14940 | 18100 | -1.214 | -0.4795 | Yes | ||
| 142 | GAB1 | 18828 | 18157 | -1.275 | -0.4636 | Yes | ||
| 143 | TNFAIP3 | 19810 | 18163 | -1.292 | -0.4446 | Yes | ||
| 144 | RHOB | 4411 | 18166 | -1.305 | -0.4253 | Yes | ||
| 145 | STAT3 | 5525 9906 | 18210 | -1.378 | -0.4072 | Yes | ||
| 146 | LCK | 15746 | 18240 | -1.451 | -0.3872 | Yes | ||
| 147 | RIT1 | 5382 | 18243 | -1.461 | -0.3656 | Yes | ||
| 148 | EHD4 | 8275 | 18265 | -1.508 | -0.3443 | Yes | ||
| 149 | SQSTM1 | 9517 | 18281 | -1.543 | -0.3221 | Yes | ||
| 150 | MAPKAPK2 | 13838 | 18287 | -1.556 | -0.2993 | Yes | ||
| 151 | KRAS | 9247 | 18298 | -1.598 | -0.2761 | Yes | ||
| 152 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 18347 | -1.737 | -0.2528 | Yes | ||
| 153 | MAPK9 | 1233 20903 1383 | 18355 | -1.758 | -0.2271 | Yes | ||
| 154 | TP53 | 20822 | 18549 | -2.941 | -0.1938 | Yes | ||
| 155 | EGR1 | 23598 | 18576 | -3.676 | -0.1405 | Yes | ||
| 156 | MAPK1 | 1642 11167 | 18579 | -3.793 | -0.0842 | Yes | ||
| 157 | RAN | 5356 9691 | 18611 | -5.795 | 0.0003 | Yes |
-MEDIATED_SIGNALING_106.png)